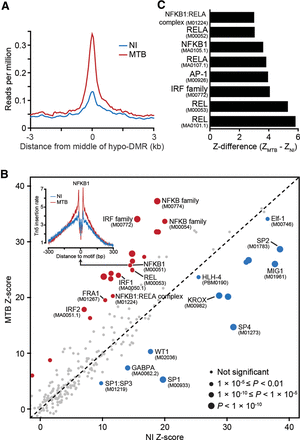
MTB-DMRs are bound by signal-dependent transcription factors. (A) Tn5-accessibility profiles before (NI) and after MTB infection, ±3 kb around the midpoints of hypomethylated regions. (B) Scatterplot comparing transcription factor occupancy score predictions between noninfected (x-axis) and MTB-infected DCs (y-axis). The size of the dots is proportional to the level of statistical significance supporting differential binding in response to MTB infection. Red dots represent TFs that show evidence for increased binding after MTB infection, and blue dots represent TFs that show evidence for decreased binding after infection. The inset on the top left corner shows the genome-wide footprint of the NF-κB (p50) motif (motif ID: M00051) in noninfected (blue) and MTB-infected DCs (red). In this example, the footprint in MTB-infected DCs is clearly stronger, which supports increased TF binding to the NF-κB (p50) motif genome-wide, upon MTB infection. (C) TF motifs (motif IDs in parentheses) that show significantly increased binding in hypomethylated regions after MTB infection.











