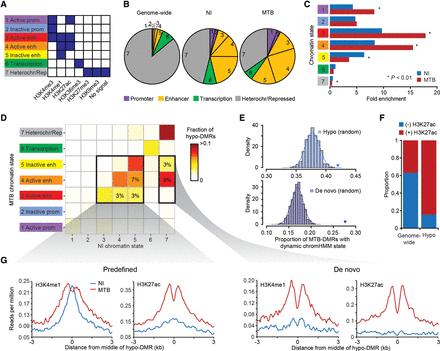
MTB-DMRs overlap with enhancer elements that become active upon infection in hypomethylated regions. (A) Combination of histone patterns used to define the seven chromatin states. The precise relative contribution of each chromatin mark to each of the chromHMM-defined states can be found in Supplemental Figure S3. Note that state 7 was defined by either no signal or the presence of either H3K27me3/H3K9me3. (B) Pie charts showing the distribution of chromatin state annotations genome-wide (on noninfected cells) and within all MTB-DMRs in either noninfected (NI) or MTB-infected cells. The chromatin state codes are as defined in A. (C) Fold enrichments of the different chromatin states within hypomethylated regions as compared to genome-wide expectations in noninfected (blue) and MTB-infected cells (red). (D) Heat map of the proportion of hypomethylated regions by chromatin transition state. The x-axis represents the chromatin states defined in noninfected DCs and the y-axis the chromatin state of the same region in MTB-infected DCs. The two bold inner boxes indicate two subgroups of hypomethylated regions, (left) predefined enhancers (detectable enhancers in noninfected DCs) and (right) de novo enhancers (detectable enhancers only in MTB-infected DCs). The numbers inside the cells refer to the proportion of hypomethylated regions that undergo each of the highlighted transitions. (E) (Top panel) Histogram showing the observed proportion of regions that change chromatin state after infection (any transition) when sampling 1000 random sets of regions matched to the chromatin states found in noninfected samples within hypomethylated regions. Each random set contains the same number of hypomethylated regions as those identified in the true data (n = 1714). The blue triangle represents the observed proportion of hypomethylated regions that changed chromatin state in response to MTB infection. (Bottom panel) Same as above but focusing on regions of the genome labeled as heterochromatin/repressed before infection (state 7; n = 790) that gain de novo enhancer marks upon MTB infection (states 3, 4, or 5). The purple triangle represents the proportion observed within the true set of hypomethylated regions. (F) Bar plot showing the proportion of hypomethylated regions that overlap with enhancers and show dynamic changes in chromatin state, as defined by the gain or loss of H3K27ac mark. (G) Composite plots of patterns of H3K4me1 and H3K27ac ChIP-seq signals ±3 kb around the midpoints of hypomethylated regions (x-axis) overlapping with predefined (left) and de novo (right) enhancers.











