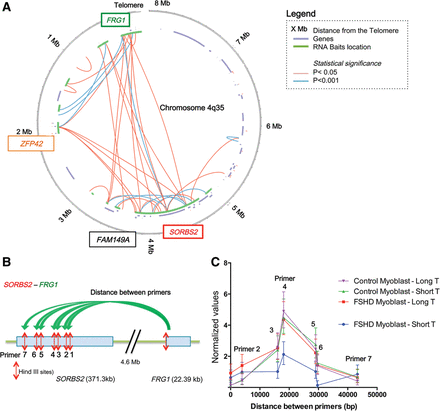
Long-range interactions revealed by locus-specific Hi-C at the 4q35 locus. (A) Circos plot (Krzywinski et al. 2009) of the 3D organization of the 4q35.2 locus in FSHD myoblasts with long telomeres after Hi-C 4q35-specific experiment. Data were corrected and normalized to background using HOMER software. Only true significant interactions are shown (red: P < 0.05; blue: P < 0.001). The last 8 Mb of Chromosome 4q are represented as a circle, the location of known genes are shown as segments in a purple inner circle, and the location of biotinylated bait sequences is shown in green. The locations of the probes used for chromosome conformation capture (3C) and in situ hybridization are indicated in colored boxes (green for FRG1 and red for SORBS2). (B) Schematic representation of the Hi-C validation by 3C for a 4.6-Mb loop between FRG1 and SORBS2. A forward primer adjacent to a HindIII site in the FRG1 sequence was used with a variety of reverse primers adjacent to HindIII sites in the SORBS2 sequence. Following crosslinking, digestion with HindIII, ligation, and crosslink reversion, DNA was amplified with the different primer pairs. Each assay was done with droplet digital PCR (ddPCR) and normalized to a 3C assay performed in a control region. (C) Graphical representation of 3C quantification of the FRG1-SORBS2 loop in control and FSHD myoblasts with long and short telomeres. Only FSHD myoblasts with short telomeres differ, with a decrease of 48.5% in the signal corresponding to the center of interaction. Assays were done in triplicate and measured in duplicate (Mean ± SD shown for each primer pair).











