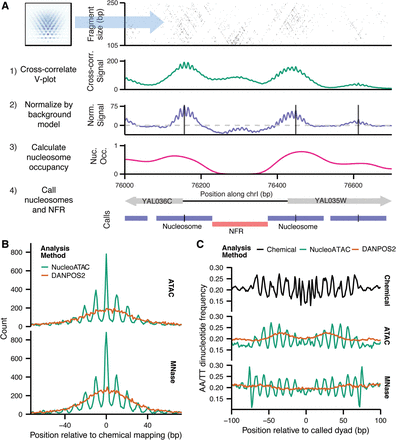
NucleoATAC enables high-resolution nucleosome positioning. (A) Schematic of NucleoATAC workflow. First, the V-plot nucleosome signature is cross-correlated against a 2D fragment size versus fragment midpoint representation of ATAC-seq data at a locus. The signal is then normalized by a background model (based on sequence bias and read depth) to obtain a normalized signal. Nucleosome occupancy is calculated using the local fraction of nucleosomal fragments. The normalized cross-correlation signal and nucleosome occupancy tracks are used to assign nucleosome and nucleosome-free (NFR) positions. (B) Distance of dyad calls from different assays (ATAC on top panel; MNase on bottom) using either NucleoATAC (green) or DANPOS2 (orange). (C) AA/TT dinucleotide pattern around nucleosome dyad calls determined by chemical mapping (top panel), or from ATAC-seq (middle panel), or MNase-seq (bottom panel) using either NucleoATAC (green) or DANPOS2 (orange).











