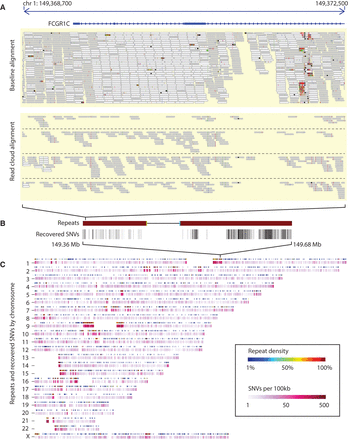
Whole-genome SNV calling on the IDC sample. (A) Comparison of the initial baseline short-read alignments of all the wells merged together with four wells aligned with RFA (from two distinct haplotypes), in a region overlapping the FCGR1C gene. (B) Placement of recovered SNVs within the surrounding 300-kbp region. (C) Density of recovered SNVs throughout the whole genome (bottom track), by chromosome, compared to density of segmental duplications (top track). Long clustered regions of recovered SNVs coincide with dense regions of annotated segmental duplications.











