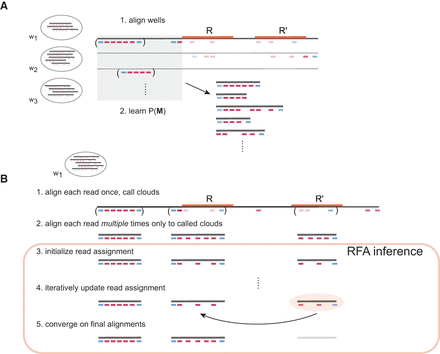
RFA overview. (A) Wells wn from the sample are first aligned to the reference using an existing short-read aligner, and uniquely mapped read clouds are used to learn a prior P(M), which captures protocol properties such as the long fragment size distribution. (B) Each well is aligned separately with the aid of a short-read aligner to determine candidate source long fragment locations as well as multiple candidate short-read alignments to the long fragments. Finally, MAP inference is performed to converge on optimal alignments. In this example, RFA successfully determines the correct repeat copy R that overlaps with a source long fragment.











