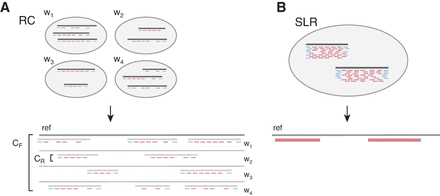
Read clouds (RC) and synthetic long reads (SLR) obtained by Illumina TruSeq Synthetic Long-Read sequencing. Each well initially contains long molecules that represent a small fraction of the target genome; reads from each long molecule are separated in genomic coordinates within the target genome, and therefore, clusters of such reads (read clouds) are formed with each cluster originating from one source fragment. Blue reads denote end-markers of the source fragments and may not always be present as sequenced short reads. (A) In the RC approach, long fragments from several wells wn are sequenced to a shallow depth and aligned to the reference to obtain read clouds. Pooling of reads across several read clouds allows inference of the variation in the underlying long fragments. (B) In the SLR approach, long fragments are sequenced to a much higher depth to enable de novo assembly of synthetic long reads. For the same total sequencing budget C, the RC approach covers proportionally more target genome space than the SLR approach.











