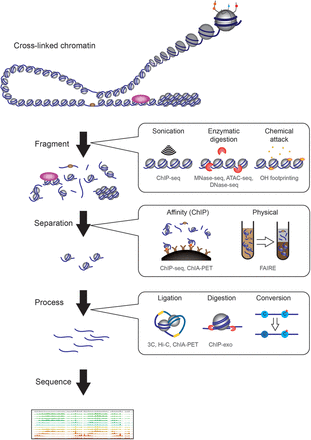
There are many broad classes of methods used in epigenomic studies prior to sequencing. We can roughly categorize methods according to three criteria: fragmentation, separation, and further processing. To interrogate chromatin, it has to be fragmented, with three broad classes of fragmentation method. First, physical fractionation methods, such as sonication, apply force to break chromatin. Second, nuclease-susceptibility methods such as MNase-seq, DNase-seq, and ATAC-seq separate regions of the genome based on their susceptibility to enzymatic attack, and typically separate “open” and accessible regions from more compact, often repressed, regions of the genome. Third, chemical susceptibility of chromatin is used in several different epigenomic methods, such as assays based on hydroxyl radical cleavage of the DNA backbone. After fragmentation, separation, either via affinity-based methods such as ChIP or physical methods such as solubility, can be used to enrich specific classes of chromatin. Finally, in some cases, further processing is used to reveal specific structural aspects of chromatin. Such methods include the proximity ligation used in the 3C family of techniques for assaying higher-order chromosome folding, or chemical conversion to reveal nucleotide modifications or precise DNA-protein crosslinks. Below each method, we list some of the assays that employ this method in gray text (e.g., ChIP, MNase-seq, etc.). Note that we do not broadly cover assays of cytosine methylation in this review, except insofar as access to methyltransferases is used as an assay for chromatin structure.











