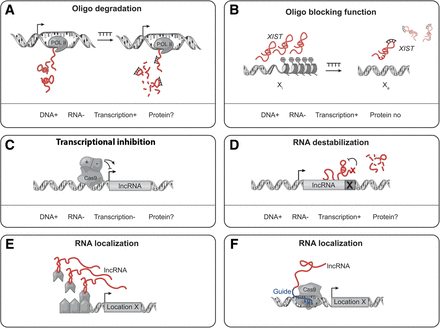
Experimental approaches to manipulate the expression, perturb the activity, or evaluate the functions of long noncoding RNAs. Most commonly used are transient expression of exogenous oligonucleotides designed to exploit the endogenous RNAi machinery or RNase H activity to degrade an RNA (A) or occlude putative functional regions (B). Recently, strategies have used the flexible CRISPR/Cas9 system to positively or negatively affect the transcription of a lncRNA gene (C) or the incorporation of an auto-catalytic regulatory RNA element to destabilize the nascent lncRNA transcript (D). An alternative to manipulating lncRNA expression levels involves the recruitment of the RNA of interest to a particular genomic locus or reporter gene through fusion of DNA-binding proteins with RNA-binding elements such as MS2 stem–loops (E) or covalent tethering of a lncRNA transcript to the CRISPR/Cas9 guide RNA (F).











