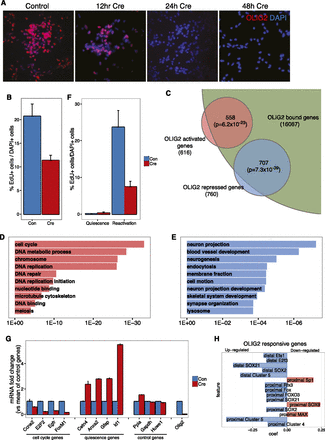
OLIG2 is a multifunctional regulator of NS cell self-renewal. (A) Immunostaining shows the rapid and complete loss of OLIG2 protein from Ollig2-conditional mutant NS cells following administration of Cre. (B) Reduced proliferation of Ollig2-mutant NS cells 48 h after Cre delivery, as measured by 3 h exposure to EdU (P-value < 0.008 Wilcoxon test). (C) Venn diagram showing the large and significant (hypergeometric test) proportion of genes deregulated in Ollig2-deleted cells that are associated with OLIG2-bound DHSs. (D,E) GO biological processes enriched among the genes down- (D) and up-regulated (E) by Ollig2-deletion. DAVID P-values are shown. (F) Ollig2-mutant cells normally arrest their proliferation when treated with quiescence-inducing medium, but are less likely to re-enter the cell cycle when returned to self-renewal medium, as measured by 3 h of EdU exposure (P-value < 0.008, Wilcoxon test). (G) Reduced expression of cell cycle regulators and active NS cell marker EGFR and inappropriate expression of quiescence-associated genes in mutant cells that have been stimulated to resume proliferation, as measured by quantitative RT-PCR. (H) Top 15 model coefficients predictive of genes up- (blue) or down-regulated (red) in the models predicting genes responsive to elimination of OLIG2 protein.











