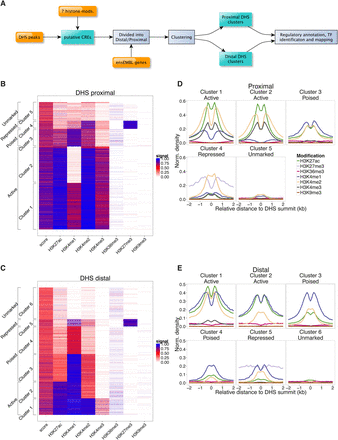
Clustering analysis reveals multiple classes of epigenetically distinct gene regulatory regions in NS cells. (A) Schematic depicting the classification and regulatory annotation of DHS regions employed in this study. (B,C) Heatmaps of TSS-proximal (B) and TSS-distal (C) DHS regions generated by k-means clustering analysis according to the local ChIP-seq signal for seven histone modifications (x-axis). Five distinct proximal clusters and six distal clusters were identified. Regulatory designations (y-axis) are given according to our downstream analyses, as described in the Results section. (D,E) Aggregate plots of histone modification signal centered upon the point of maximal DNase-seq enrichment for each proximal (D) and distal (E) putative regulatory element.











