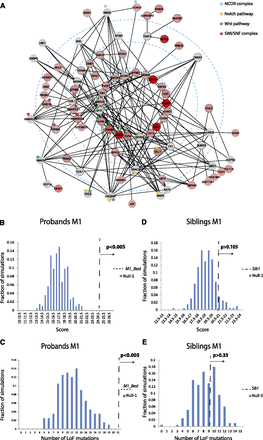
Module M1. (A) Genes detected as part of module M1_Extended are displayed as graph nodes using Cytoscape (Shannon et al. 2003). Node colors reflect the score of each gene based on the number and type of de novo mutations: The more intense red color
indicates a higher score, while gray indicates a score of zero (no de novo mutations observed). Edges (black lines) between
two nodes represent genes that interact with each other according to the PPI network and are also highly coexpressed (Pearson
correlation coefficient 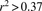 , i.e., the genes are included in the top 5% of gene pair coexpression during brain development). The innermost circle contains genes detected in > 99% of the suboptimal solutions. Subsequent concentric circles display genes found in
> 80%, 20%, and 5% (M1_Extended) of the suboptimal solutions, respectively (see Methods). Nodes with black outlines are the ones detected in the optimal
module detected (M1_Best). For a force-directed layout of this module, see Supplemental Figure 38. (B) The M1_Best score (dashed black line) shown in comparison to the top-scored module of 200 simulations using null model Null-1. (C) The number of LoF mutations covered by the top-scoring module (M1_Best) found using proband mutations versus the number of simulated LoF mutations covered by the top-scoring modules found under
the same simulation. (D) The score of the top module found using siblings and control mutations (dashed black line) in comparison to the top-scored
module of 200 simulations with the same number of mutations using Null-1. The siblings’ simulations were performed without
using the ESP constraint (although similar results were obtained when an ESP constraint was applied). (E) The number of LoF mutations covered by the top-scoring module found using siblings’ mutations versus the number of simulated
LoF mutations covered by the top-scoring modules found using Null-1. Sibling simulations were performed without filtering
based on ESP controls.
, i.e., the genes are included in the top 5% of gene pair coexpression during brain development). The innermost circle contains genes detected in > 99% of the suboptimal solutions. Subsequent concentric circles display genes found in
> 80%, 20%, and 5% (M1_Extended) of the suboptimal solutions, respectively (see Methods). Nodes with black outlines are the ones detected in the optimal
module detected (M1_Best). For a force-directed layout of this module, see Supplemental Figure 38. (B) The M1_Best score (dashed black line) shown in comparison to the top-scored module of 200 simulations using null model Null-1. (C) The number of LoF mutations covered by the top-scoring module (M1_Best) found using proband mutations versus the number of simulated LoF mutations covered by the top-scoring modules found under
the same simulation. (D) The score of the top module found using siblings and control mutations (dashed black line) in comparison to the top-scored
module of 200 simulations with the same number of mutations using Null-1. The siblings’ simulations were performed without
using the ESP constraint (although similar results were obtained when an ESP constraint was applied). (E) The number of LoF mutations covered by the top-scoring module found using siblings’ mutations versus the number of simulated
LoF mutations covered by the top-scoring modules found using Null-1. Sibling simulations were performed without filtering
based on ESP controls.











