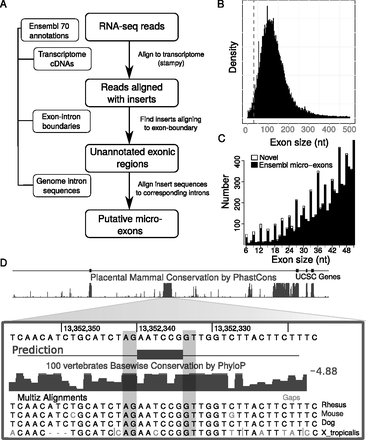
Identification of novel micro-exons. (A) Flowchart of our micro-exon discovery pipeline. Ensembl release 70 annotation was first used to build all cDNA transcripts on which RNA-seq reads were mapped using Stampy (Lunter and Goodson 2011). Reads aligning with insertions of up to 51 nt in length were then scanned to identify those whose insertions aligned to exon-exon boundaries. Subsequently, the inserted sequences were aligned to the intronic sequences separating the corresponding exons. Putative novel micro-exons were then defined as exons that were flanked by canonical splice sites and were supported in at least 15% of all samples. (B) The density of internal exon sizes shows that the majority is distributed around 140 nt in length, while there is a sharp decrease in the number of exons shorter than 51 nt (dashed line) as exon size decreases. (C) Previously annotated micro-exons from Ensembl release 70 that show evidence for expression in brain samples (black) compared to novel predicted micro-exons expressed in brain samples (white). Although the annotation of internal exons of sizes 22–51 nt appears to be nearly complete, we identified a large number of novel micro-exons between 6 and 21 nt in length. (D) Example of a novel predicted micro-exon. This micro-exon is only 6 nt in length and lies within a conserved region of the CACNA1 gene. The splice sites of this micro-exon are conserved in mammals and in Xenopus.











