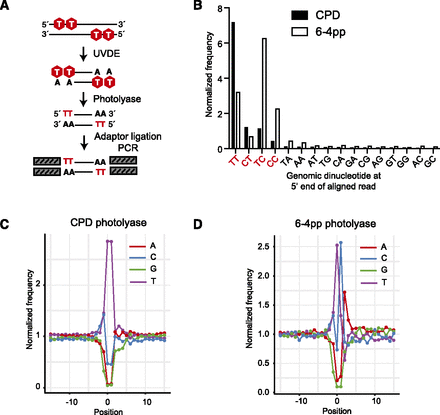
Excision-seq mapping of dipyrimidines in the budding yeast genome. (A) In predigestion Excision-seq for pyrimidine dimers, UV-damaged DNA is cleaved with UVDE, releasing double-stranded fragments with five dipyrimidines (red). Fragments are treated with CPD or 6-4pp photolyase enzymes, repairing dipyrimidines to “mono” pyrimidines, and yielding ends compatible with polishing, ligation, and PCR. (B) Analysis of sequencing libraries treated with CPD or 6-4pp photolyase prepared from UV-irradiated DNA showed an enrichment of sequence reads with dipyrimidine ends (red text) relative to genomic dinucleotide content and recapitulated the known specificity of the photolyase enzymes (Chowdhury and Guengerich 2008). (C) Frequency of nucleotides relative to mapped positions of sequences from predigestion Excision-seq libraries for mapping cyclobutane dimers in S. cerevisiae. Position 0 corresponds to the mapped position of the 5′ end of 5,063,196 sequencing reads. (D) Frequency of nucleotides relative to mapped positions of sequences from predigestion Excision-seq libraries for mapping 6-4 photoproducts in S. cerevisiae. Position 0 corresponds to the mapped position of the 5′ end of 3,655,251 sequencing reads.











