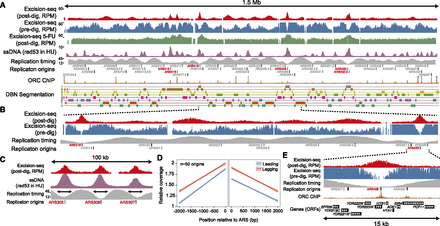
Excision-seq maps of uracil content in the budding yeast genome. (A) Data showing the entire yeast chromosome 4 for dut1-1 ung1∆ yeast using post-digestion Excision-seq (red, reads per million [RPM]), predigestion Excision-seq (blue, RPM), post-digestion Excision-seq data for ung1∆ yeast treated with 5-fluorouracil (5-FU) (green, RPM), single-stranded DNA accumulation caused by hydroxyurea treatment of a rad53 yeast strain (purple, arbitrary units) (Feng et al. 2006), replication timing data (Trep, minutes replicated after G1 release) (Raghuraman et al. 2001) (gray), annotated origins of replication (Nieduszynski et al. 2007), ORC chromatin immunoprecipitation signals (Eaton et al. 2010) (brown, coverage), and labeled segments from an eight-state DBN segmentation (Hoffman et al. 2012) incorporating replication timing (Yabuki et al. 2002) and post-digestion Excision-seq mapping of uracil. (B) A 450-kb region of chromosome 4 highlights patterns of uracil incorporation in early-replicating origins (ARS418 and ARS428), as well as uracil depletion in late-replicating regions. (C) Correspondence of peak widths between post-digestion Excision-seq (red) and ssDNA accumulation (Feng et al. 2006) (purple) at three early-replicating origins in a 100-kb region of chromosome 3. (D) Post-digestion Excision-seq measurement of uracil content for 50 early-replicating origins. Lagging strands have ∼1.3-fold higher relative coverage than leading strands in post-digestion Excision-seq data, reflecting increased uracil content in leading strands. (E) A 15-kb region of chromosome 4 highlights patterns of uracil incorporation at the early-replicating origin ARS428.











