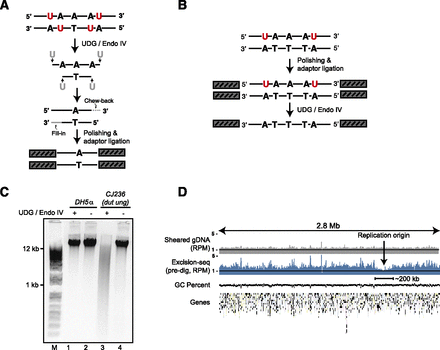
Excision-seq methods for mapping modified nucleobases in genomic DNA. (A) In “predigestion” Excision-seq for uracil, uracil-containing DNA is cut with a base excision repair enzyme (e.g., E. coli UDG, red). Released fragments are end-repaired, A-tailed, ligated to adaptors, and PCR amplified. Sequences derived from this library identify the positions of modified bases (e.g., one base upstream of the 5′-most position of the read). (B) In “post-digestion” Excision-seq for uracil, DNA is sheared mechanically, then treated by standard polishing and ligation. A base excision enzyme cleaves one or both strands containing modified bases. Intact strands remaining after digestion are PCR amplified and sequenced. (C) Genomic DNA isolated from dut ung E. coli is digested by UDG and T4 endonuclease IV (cf. lanes 3 and 4), whereas genomic DNA isolated from a wild-type strain is not digested (lanes 1 and 2). (D) Normalized coverage from shotgun sequencing of mechanically sheared genomic DNA (gray, reads per million [RPM]) and predigestion Excision-seq for uracil (blue, RPM) for a 2.8-Mb region of the E. coli chromosome. GC-content and the positions of protein-coding genes are plotted below. Uracil content is lowest in a region centered on the origin of replication, encompassing ∼200 kb of DNA.











