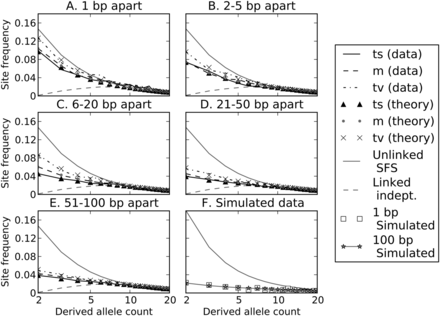
Site frequency spectra of perfect LD mutations. Each of panels A through E contains site frequency spectra of transitions, mixed pairs, and transversions found in perfect LD in the 1000 Genomes data. Singletons are excluded because they cannot be phased and therefore perfect LD status cannot be determined. For comparison, each panel contains the population-wide SFS of unlinked SNPs as well as the inferred SFS of linked independent mutations. SNP pairs are binned according to the distance between them, showing that close-together SNPs and transversions have spectra closer to the population SFS, while far-apart SNPs and transitions appear more weighted toward linked independent mutations. Dotted lines show the frequency spectra predicted by Equation 1 for each length and pair type category, assuming that the gray dashed line [m (theory)] depicts the correct SFS of linked independent mutations and that Figure 7 shows the correct MNM percentages in each category. For comparison, panel F shows a population SFS and perfect LD frequency spectrum obtained from data simulated under a human demographic model. In the simulated data, there is no difference between the frequency spectra of linked independent mutations that lie 1 bp apart versus 100 bp apart.











