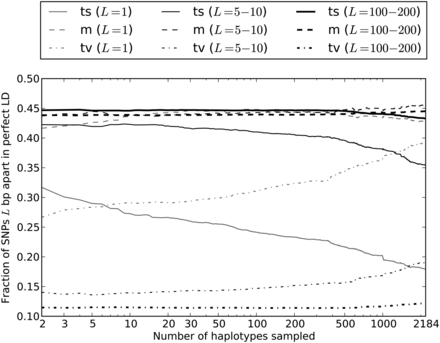
Enrichment of transversion pairs and MNMs with increasing sample size. We generated subsamples of the 1000 Genomes data containing 2–2184 haplotypes and computed the percentages of transversion pairs, transition pairs, and mixed pairs for perfect LD SNPs in each data set. As the number of sampled haplotypes increases, the percentage of perfect LD SNPs that are MNMs should increase, leading to an increase in the frequency of transversions and a decrease in the frequency of transitions. This effect is most apparent when the SNPs are adjacent (1 bp apart) or very close (5–10 bp apart). However, perfect LD SNPs that lie 100–200 bp apart display the same pattern, indicating that MNMs spanning 100–200 bp are much less common but are still evident in samples of many lineages.











