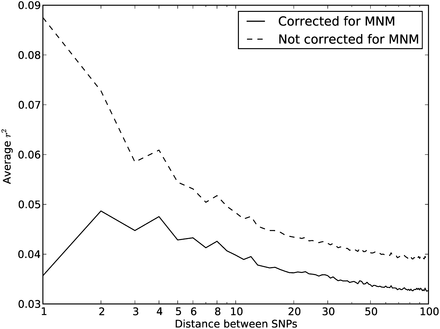
Average r2 LD correlations between 1000 Genomes SNP pairs. The correlation coefficient r2 between allele frequencies at neighboring sites is often used to measure the decay rate of genealogical correlation with
genomic distance. However, we have seen that multinucleotide mutation creates excess LD compared to the expectation under
independent mutation. We computed the average r2 across all SNP pairs L bp apart on chromosome 22, then corrected this value for the presence of MNM.  is lower at a distance of 1 bp than a distance of 2 bp because of double deaminations at CpG sites that occur on separate
lineages.
is lower at a distance of 1 bp than a distance of 2 bp because of double deaminations at CpG sites that occur on separate
lineages.











