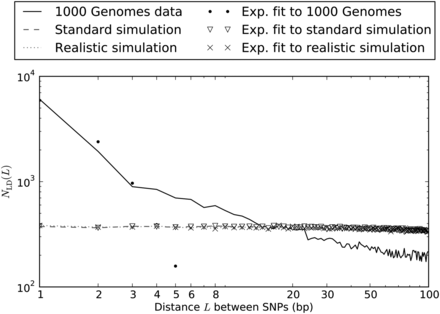
Nearby SNPs in LD: 1000 Genomes Phase I data vs. simulation under mutational independence. When we simulated 2184 haplotypes under a realistic demographic model, we observed ∼37,000 SNP pairs in LD separated by <100 bp in a sample of total length 4.8 × 108 bp. Their spacing was distributed almost uniformly between 1 and 100 bp. We observed much less uniformity in the distribution of distances between SNP pairs in LD in the 1000 Genomes data, with an extreme excess of SNPs in LD at 1–2 bp and a less extreme excess of SNPs at distances up to 20 bp apart. (Note that the axes are logarithmically scaled, making exponential curves appear concave downward.)











