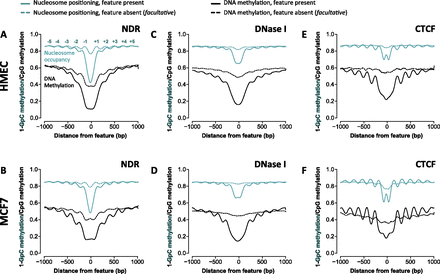
NOMe-seq reveals that nucleosomes are organized throughout the genome, at nucleosome-depleted regions (NDRs) and facultativeNDRs (fNDRs). (A–F) Each genomic feature (NDR, DNase hypersensitivity, or CTCF binding site) was called as “present” (solid lines) or “absent” (broken lines) in HMEC (upper panels) or MCF7 (lower panels). Nucleosome occupancy and endogenous methylation (CpG) were mapped ±1000 bp from the center (“0”) of each NDR, DNase I hypersensitive site, or CTCF site. (A,B) NOMe-seq demonstrates that nucleosomes are present on either side of a NDR (solid teal lines) and across the genome, even at less pronounced or absent NDRs (broken teal line). DNA methylation is phased alongside nucleosomes (black lines) regardless of nucleosome occupancy; however, DNA methylation is typically low (∼20%) within NDRs (solid black line). (C,D). DNase I hypersensitive sites are characterized by organized nucleosomes (solid teal lines), whereas DNA methylation (solid black lines) is low at the center, then phased, and then increases with distance from the center of the DNase I site (solid black line) similar to patterns observed around NDRs. (E,F). CTCF is associated with nucleosome patterns (solid teal lines) and phased peaks of DNA methylation (solid black lines) between the nucleosomes. In the absence of CTCF, nucleosomes are not phased (broken teal lines), and a continuous high level of DNA methylation is observed (∼60%).











