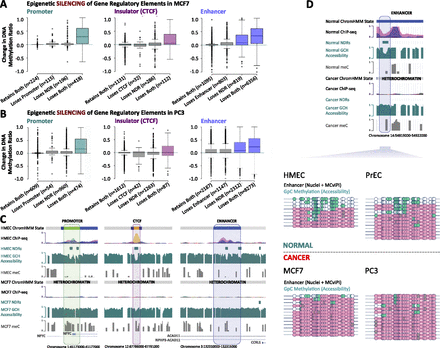
Epigenetic silencing of enhancers and insulators, in addition to promoters, in cancer cells. (A) Promoters (broadly, nucleosome-depleted, and H3K4me3 enriched) were defined in HMEC cells. These exact genomic regions were subject to ChromHMM classification in MCF7 cells and the extent and type of epigenetic silencing (nucleosome acquisition, loss of active epigenetic marks, and gain of DNA methylation) was determined at promoters (left panel, teal). Similarly, insulators (broadly, nucleosome-depleted, and CTCF; center panel, purple) and enhancers (broadly, nucleosome-depleted, and H3K4me1 enriched; right panel, blue) were defined in HMEC cells. These exact genomic regions were subject to ChromHMM classification in MCF7 cells, and the extent and type of epigenetic silencing was determined at insulators and enhancers. (B) As in A, for PrEC and PC3. (C) Screenshots showing epigenetic reprogramming at a promoter (left panel), a CTCF site (middle panel), and an enhancer (right panel). For ChIP-seq tracks: (orange) CTCF; (pink) H3K27ac; (purple) H3K4me1; (green) H3K4me3; and (red) H3K27me3. (D) Validation of enhancer epigenetic silencing. Representative colors for ChIP-seq tracks are as shown in C. All four cell types were treated with M.CviPI GpC methyltransferase and subjected to bisulfite conversion and cloning. Horizontal lines represent individual enhancers. Circles represent GpC dinucleotides: (white) unmethylated and inaccessible to M.CviPI; (teal) methylated and accessible to M.CviPI. Pink bars represent sites associated with nucleosomes. Regions accessible to M.CviPI (teal) indicate NDRs. Diagram is drawn to scale.











