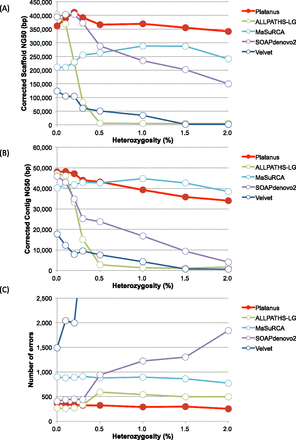
Figure 3.
Results of the benchmarks of heterozygosity simulations (C. elegans). (A) Corrected scaffold-NG50 calculated by GAGE. (B) Corrected contig-NG50. (C) Number of errors reported by GAGE. Errors are defined as inversion, relocation, or translocation.











