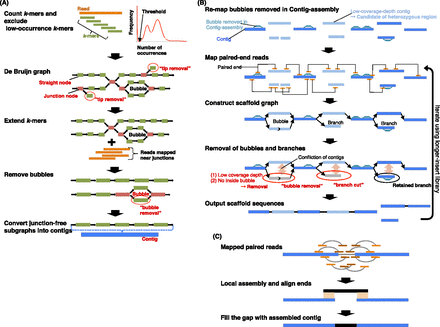
Schematic overview of the Platanus algorithm. (A) In Contig-assembly, a de Bruijn graph is constructed from the read set. Short branches caused by errors are removed by “tip removal.” Short repeats are resolved by k-mer extension, in which previous graphs and reads are mapped to nearby k-mers at the junctions. Finally, bubble structures caused by heterozygosity or errors are removed. Subgraphs without any junctions represent contigs. (B) In Scaffolding, links between contigs are detected using paired reads. The relationship between contigs is represented by the graph. Bubbles removed in Contig-assembly are remapped on contigs and utilized for mapping of paired-end reads and detection of heterozygous contigs. Heterozygous regions are removed as bubble or branch structures on the graph by the “bubble removal” or “branch cut” step. These simplification steps are characteristic of Platanus and especially effective for assembling complex heterozygous regions. (C) In Gap-close, paired reads are mapped on scaffolds, and reads mapped at nearby gaps are collected for each gap. If a contig is expected to cover the gap and is constructed from collected reads, the gap is closed by the contig.











