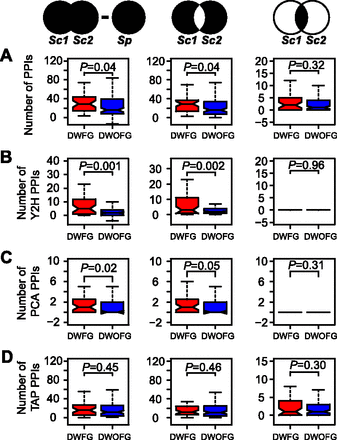
Gene duplication-associated adaptation is correlated with gains of binary protein–protein interactions (PPIs). (A) DWFG and DWOFG are compared for the difference between the total number of unique PPIs of a pair of S. cerevisiae (Sc) duplicates and that of the S. pombe (Sp) ortholog (left), the total number of unshared PPIs between a Sc duplicate pair (middle), and the number of shared PPIs between a Sc duplicate pair (right). Sc1 and Sc2 refer to the duplicate genes in S. cerevisiae, and Sp refers to their single-copy ortholog in Sp. The bottom and top of each box are the first and third quartiles, and the band inside the box shows the median. The whiskers extend to the most extreme data within 1.5 times the interquartile range below the first quartile and above the third quartile. (B–D) Same as A except that only those PPIs detected by yeast two-hybrid (Y2H) (B), protein-fragment complementation (PCA) (C), and tandem affinity purification (TAP) methods (D) are considered.











