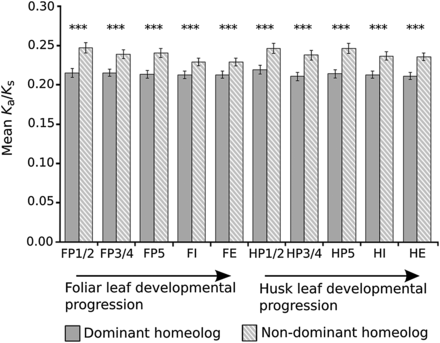
Nondominant homeolog genes experience relaxed purifying selection. Ka/Ks ratios were compared for those non-R-NF homeolog gene pairs that exhibit subgenome dominance. The mean Ka/Ks ratio of nondominant homeolog genes (those expressed at significantly lower levels than their homeolog pair) was significantly higher than the dominant homeolog (those expressed at significantly higher levels than their homeolog pair). P-values were calculated using Wilcoxon signed-rank tests. Significance levels are indicated by asterisks above the bars. (***) P < 0.001. Error bars are standard errors.











