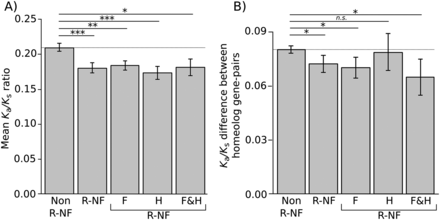
Selection acts differentially on regulatory neofunctionalized genes. (A) Mean Ka/Ks ratio of each homeolog plotted for each category. A ratio of <1 indicates negative (purifying selection), whereas >1 indicates positive selection. P-values were calculated for all R-NF homeolog gene pairs (n = 412) and each category of R-NF homeolog gene pairs using Monte Carlo resampling tests. (F) Foliar; (H) Husk; (F&H) Foliar and Husk. All R-NF categories show significantly greater purifying selection than non-R-NF homeolog gene pairs. (B) Mean magnitude of the difference in Ka/Ks ratios between homeolog pairs. The greater the difference in Ka/Ks the more selection has been relaxed on one gene in the pair. The closer the ratio is to zero the more similar the selection acting upon both homeologs. P-values were calculated using Monte Carlo resampling tests. R-NF homeolog gene pairs show a significantly smaller difference to non-R-NF homeolog gene pairs. Significance levels are indicated by asterisks above the bars. (*) P < 0.05; (**) P < 0.01; (***) P < 0.001; (n.s.) not significant. Error bars are standard errors.











