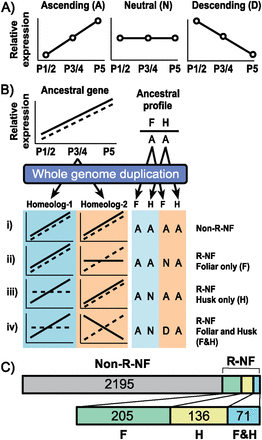
Identification and quantification of regulatory neofunctionalized genes. (A) Genes expressed in foliar (F) and husk (H) leaves were allocated to one of three expression profiles, ascending (A), neutral (N), and descending (D), based on the relative expression levels at three stages of early leaf development: plastochron (P) 1/2, P3/4, and P5 (see Methods for an explanation of profile classification). (B) Cartoon of how an example ancestral gene, with expression allocated to profile A in both F and H, could be manifested after whole-genome duplication. Blue background refers to the homeolog from subgenome-1, and orange to the homeolog from subgenome-2. Continuous line refers to the expression profile (A, N, or D) in F, whereas a dashed line refers to the expression profile in H. Four-letter expression code is made up of homeolog-1 F expression profile, homeolog-2 F expression profile, homeolog-1 H expression profile, homeolog-2 H expression profile. If the expression profile of homeolog-1 matches the expression profile of homeolog-2 in F, and the expression profile of homeolog-1 matches the expression profile of homeolog-2 in H, the gene pair is assumed to have undergone zero expression changes since duplication, and is thus nonregulatory neofunctionalized (non-R-NF). If the expression profile of homeolog-1 matches the expression profile of homeolog-2 in H, but the expression profiles in F are not the same, assuming only one expression change has occurred, this change must have been in F, and thus the gene pair is said to have been regulatory neofunctionalized (R-NF) in F only. If the expression profile of homeolog-1 matches the expression profile of homeolog-2 in F, but the expression profiles in H are not the same, assuming only one change has occurred, this change must have been in H, and thus the gene pair is said to have been R-NF in H only. If the expression profile of homeolog-1 does not match that of homeolog-2 in F, and if the expression profile of homeolog-1 does not match that of homeolog-2 in H, the gene pair is assumed to have undergone at least two changes in expression, and is thus R-NF in both F and H. (C) Results of dividing homeolog gene pairs using the method described in B. Numbers refer to homeolog gene pairs that have expression profiles consistent with the labeled category. A total of 2607 homeolog gene pairs are expressed in both foliar and husk leaf samples.











