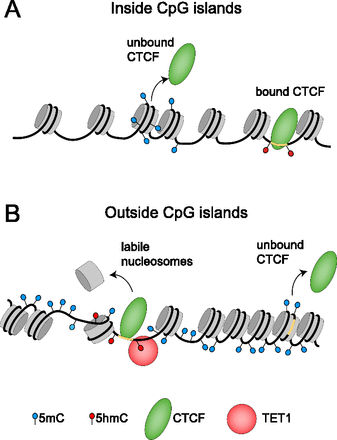
Model rationalizing the linkages between 5mC, 5hmC, TET1-, and CTCF-binding with nucleosome positioning inside and outside of CpG islands. (A) Inside CpG islands, most CpGs are unmethylated and have low nucleosome occupancy. The small fraction of CpGs in CGIs that are methylated has a nucleosome positioned preferably within the DNA methylation sites. CTCF binding inside CGIs is mostly invariant and determined by the DNA sequence: Strong constitutive CTCF sites stay unmethylated and bound by CTCF and its cobinders during the cell development, while weak sites in these regions are mostly not bound by CTCF in both ESCs and differentiated cells. (B) Outside of CGIs, the genomic DNA is mostly methylated at CpGs. In relation to nucleosome positioning, the following features were found: DNA methylation density is lowest in the middle of the nucleosome, smoothly increases toward the nucleosome entry/exit, and reaches a maximum between nucleosomes. In these regions, TET1 binding creates MNase-sensitive labile nucleosomes, which are being removed/translocated during the process of 5mC to 5hmC conversion. Variable CTCF sites are found preferentially outside CGIs, where active, TET1-dependent hydroxymethylation and associated nucleosome repositioning promotes CTCF binding. As CpGs in these regions change to a methylated state during stem cell differentiation, the formation of stably bound nucleosomes leads to a loss of CTCF at these sites.











