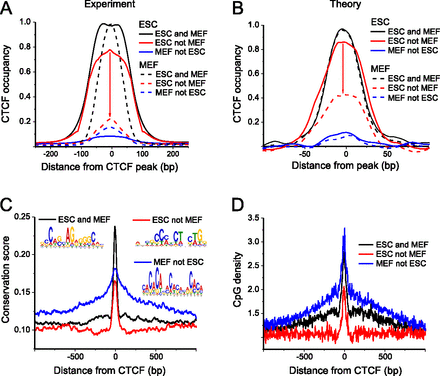
Quantitative model to predict CTCF occupancy changes due to competition with nucleosomes. (A) Experimentally observed occupancy of CTCF binding sites in ESCs and MEFs derived from ChIP-seq peak heights (Shen et al. 2012). Three subsets of CTCF sites can be distinguished: constitutive sites bound in both cell types (ESC and MEF), variable sites predominantly bound by CTCF in ESCs (ESC not MEF), and weak sites which are slightly more strongly bound in differentiated cells (MEF not ESC). (B) Predicted CTCF binding site occupancy in ESCs and MEFs when accounting for nucleosome positioning for the three binding site classes shown in panel A. (C) Conservation score of CTCF DNA binding sequence motifs for constitutive, variable, and weak binding sites. (D) Average CpG density at CTCF sites. The variable sites in the “ESC not MEF” class are located preferentially outside of CGIs.











