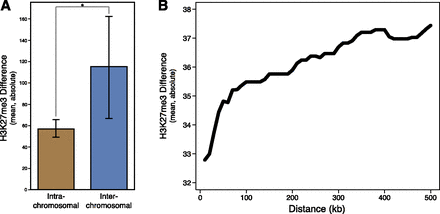
H3K27me3 changes in paralogs differ depending on duplicate location. (A) Within Drosophila simulans, interchromosomal translocations have significantly greater H3K27me3 changes (in terms of absolute value; permutation test, P < 10−5). To accurately identify chromosomal translocations, we limited comparison to the major chromosome arms (2R, 2L, 3R, 3L, and X), and considered any gene that moved between chromosome arms an interchromosomal translocation. (B) On the same chromosome, duplicated gene orthologs’ H3K27me3 changes increase with distance from the parent gene. On the y-axis, the cumulative mean change in H3K27me3 signal as a function of distance (on the x-axis). There is a significant correlation between distance and magnitude of H3K27me3 change (SCC = 0.103, P = 0.002).











