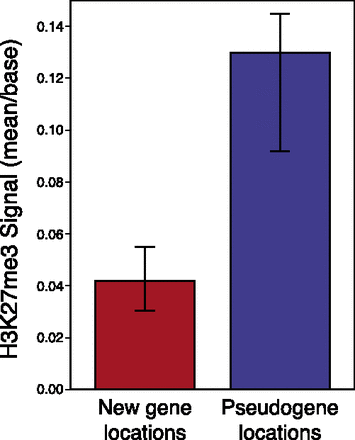
Locations into which duplicates move affects eventual pseudogenization fate. Regions in the D. simulans genome into which new genes and pseudogenes had moved in D. melanogaster show differences in mean H3K27me3 signal. On the left, functional new genes’ duplication locations show significantly less H3K27me3 signal than pseudogenes’ equivalent locations. Bars represent bootstrapped 95% confidence intervals. Note that for the purposes of this analysis, we assume that extant D. simulans H3K27me3 signal is representative of the H3K27me3 signal occurring in the ancestor of D. melanogaster in which new gene duplications occurred (see Methods).











