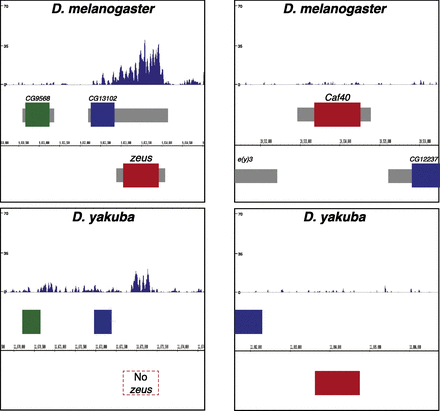
The novel gene duplicate Zeus has undergone rapid gain of H3K27me3. (Left) A Genome Browser snapshot indicating H3K27me3 signal at the Zeus (Rcd-1r) locus in D. melanogaster. The first window indicates the normalized, input-subtracted H3K27me3 signal pooled from two separate biological replicates in D. melanogaster. (Below) Boxes indicate protein-coding genes: CG9568 (green), CG13102 (blue), and Zeus (red). The next window is as above, but in an outgroup species (D. yakuba), which does not possess the Zeus retroposition. Browser snapshots are aligned such that orthologous genes match position. Note that the H3K27me3 level around Zeus is substantially higher in D. melanogaster relative to the equivalent region in D. yakuba. (Right) As above, but focusing instead on the parental gene CAF40 (Rcd-1; red), flanked on the left by e(y)3 (gray), and on the right by CG12237 (blue). Note that CAF40 possesses little to no H3K27me3 signal, in strong contrast to Zeus. The sequence identity between Zeus and CAF40 is 71% (Quezada-Díaz et al. 2010).











