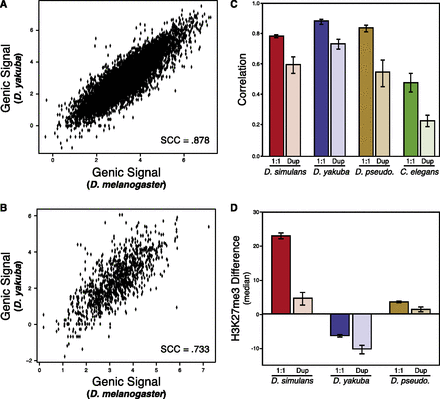
Duplicated genes have less conserved H3K27me3 signal. (A) As in Figure 1A, single-copy orthologous gene H3K27me3 conservation between melanogaster and yakuba. (B) As in A and Figure 1A, but depicting duplicated gene orthologs. (C) Overall pattern of H3K27me3 conservation in different gene sets. Each pair of bars represents one species comparison; bars on the left are the correlation coefficient of single-copy orthologs; on the right, the correlation coefficient of duplicated gene orthologs. Each bar represents a single pairwise comparison’s Spearman rho, compared against Drosophila melanogaster: (red) simulans; (purple) yakuba; (brown) pseudoobscura; (green) C. elegans. Overall correlation decreases in each case. Lines are 95% bootstrapped confidence intervals for each correlation; we account for the differences in sample size (single-copy orthologs are more common than duplicated gene orthologs) by resampling the number of duplicated gene orthologs in each case. (D) Duplicated gene orthologs are more likely to lose H3K27me3. Each pair of barplots is one species (simulans, yakuba, pseudoobscura); the left in each pair is the median H3K27me3 change in duplicated genes relative to the single gene in the other genome; while the right is the median in single-copy orthologs. Each pair shows that duplicated genes are biased toward the loss of H3K27me3 signal. For each pair (by permutation test): simulans, P < 10−5; yakuba, P < 10−5; and pseudoobscura, P < 10−5.











