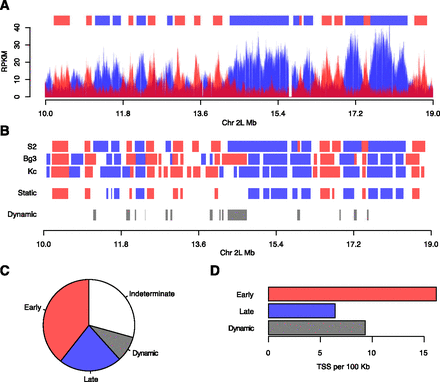
Identification of static and dynamic replication domains. (A) Segmentation of the Drosophila genome into discrete early and late replicating domains. A three-state hidden Markov model representing early, late, or indeterminate replication timing was used to define early (red) and late (blue) replicating domains (top) from the RPKM data from the early and late S-phase fractions. A 9-Mb portion of chromosome 2L is depicted for S2 cells. (B) Identification of static and dynamic replication domains. The early and late replication domains were compared between cell lines. Domains that maintain their replication timing in all cell lines (static) or those that change their replication timing in at least one cell line (dynamic) were identified. (C) Pie chart representing the fraction of static early (red, 39.3%), static late (blue, 22.2%), dynamic (gray, 8.1%), and indeterminate (30.4%, white) replication timing domains. (D) Density of TSS as a function of domain length (per 100 kb) for static early (red), static late (blue), and dynamic (gray) replication domains.











