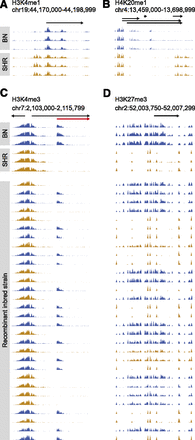
Strain-specific and allele-dependent histone methylation marks in progenitor and RI rats. Example of strain-specific H3K4me1 peaks (A), H4K20me1 peaks (B), H3K4me3 peaks (C), and H3K27me3 peaks (D) in three BN and three SHR rats. In C and D, RI strains were split according to their genotype at the position of the histone mark and are depicted in blue (BN genotype) or orange (SHR genotype). Genomic positions, Ensembl genes, and their direction of transcription are indicated by arrows. In C, strain-specific H3K4me3 marks colocalized with an alternative TSS, which was detected using RNA-seq and is depicted in red.











