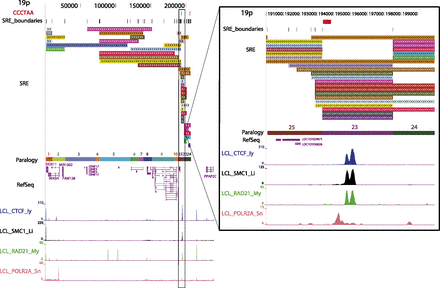
Subtelomere annotation features. The first 250 kb of the 19p subtelomere assembly is shown to illustrate key features of subtelomere sequence organization annotated on our browser. Coordinate 1 on the browser corresponds to the centromeric end of the terminal repeat tract [i.e., the last (CCCTAA)n repeat unit before subtelomere DNA starts]. The 207-kb-long SRE region on 19p is subdivided into duplication modules (“duplicons”) defined by segments of similarity (>90% nucleotide identity, >1 kb in length) between 19p and other subtelomeres (Ambrosini et al. 2007). Each rectangle represents a separate duplicon. Duplicated segments are identified by chromosome (color) as described previously (Ambrosini et al. 2007); additional details included on the live browser but omitted for the sake of clarity include the subject subtelomere identity, starting and ending coordinates of the duplicon in the subject subtelomere sequence, and the percentage of nucleotide sequence similarity of non-RepeatMasked sequences from the duplicon segment of the subject subtelomere to 19p (vader.wistar.upenn.edu/humansubtel). Each SRE boundary is indicated on a single track (SRE_boundaries), as are the internal telomere-like sequence (ITS) islands as defined in Methods (red ticks in the CCCTAA track). Gene models for transcripts included in the RefSeq (shown) (Pruitt et al. 2012) and Ensembl (hidden in this figure) (Flicek et al. 2012) transcript databases were mapped using Spidey (Wheelan et al. 2001). The paralogy track corresponds to the blocks, as shown in Figure 1. Enrichment profiles for four ChIP-seq data sets originally mapped only to subterminal DNA sequences (Deng et al. 2012) are displayed. (Inset) Close-up view of an internal SRE boundary region showing the association of the boundaries with an ITS (red rectangle on top line) and enrichment peaks for CTCF, cohesin subunits SMC1A and RAD21, and RNA polymerase II large subunit (POLR2A).











