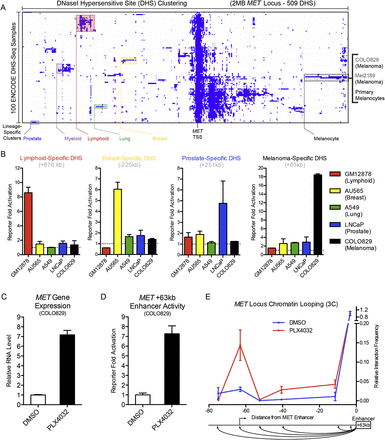
Discovery of a lineage-specific, kinase inhibitor-responsive MET enhancer. (A) DNase I hypersensitive site (DHS) clustering at the MET gene locus. Lineage-specific clusters are noted in color, with individual cell types within the melanocyte lineage enumerated at right. (B) Enhancer reporter activation levels of selected lineage-specific DHS over an empty vector background in lymphoid, breast, lung, prostate, and melanocyte cell types. (C) MET gene expression in human melanoma cell line COLO829 after a 24-h treatment with DMSO or a selective oncogenic BRAF inhibitor PLX4032. (D) Enhancer reporter activation levels of the melanoma lineage-specific MET +63-kb enhancer in COLO829 cells after a 24-h treatment with DMSO or PLX4032. (E) Chromosome conformation capture (3C) at the MET locus as a function of BRAF inhibition with the +63-kb enhancer as an anchor region. Schematic of the assayed genomic locations below.











