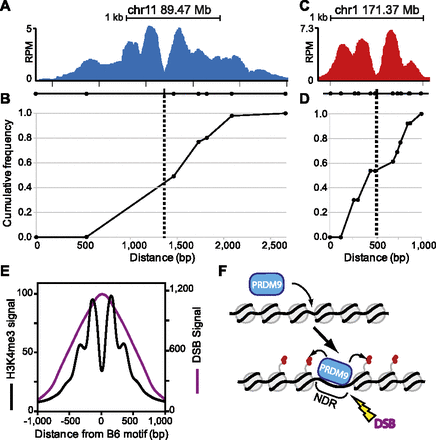
Location of meiotic crossing-over is constrained by PRDM9-dependent chromatin modification. (A) ChIP-seq coverage profile for the PRDM9Dom2-activated hotspot at chr11 89.47 Mb showing nucleosome peaks and central NDR. The black line underneath the nucleosome profile indicates the positions of SNPs (circles) between the B6 and CAST strains used to map recombination and the PRDM9Dom2 binding site from Supplemental Figure S5 (black box). (B) Cumulative frequency of recombination in B6 × CAST F1 progeny plotted against the scaled distance across the H3K4me3 signal track (n = 56 recombinant mice). (C,D) Similar to A and B except for the PRDM9Cst-activated hotspot at chr1 171.37 Mb. (C) ChIP-seq coverage profile showing nucleosome peaks and central NDR. SNPs assayed for recombination between B6 and CAST strains are indicated below as circles and PRDM9Cst binding site from Figure 4F is shown (black box). (D) Cumulative frequency of recombination at this hotspot (n = 13). (E) Aggregation plot of Prdm9Dom2 hotspots for mean H3K4me3 signal (black line) and mean DMC1 ChIP-seq signal representing the position of meiotic DSBs (purple line, n = 7235 peaks, DSB data from Brick et al. [2012]). (F) Model of PRDM9 activation of hotspots. Prior to PRDM9 binding, hotspot nucleosome positions can be variable. Upon binding, PRDM9 modifies surrounding nucleosomes (red circles), which are remodeled to create a central nucleosome-depleted region (NDR). Later, DNA double-strand breaks (DSBs) are introduced at the NDR.











