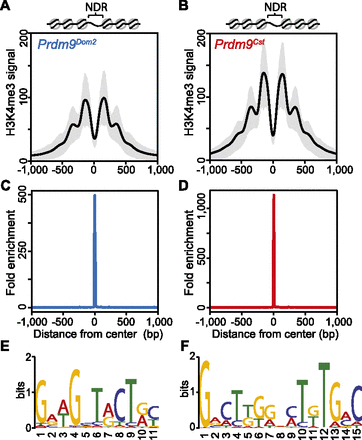
H3K4me3 nucleosomes are organized symmetrically around predicted PRDM9 binding motifs, creating a central nucleosome-depleted region (NDR). (A) Aggregate plot of H3K4me3 signal across a 2-kb window from PRDM9Dom2-dependent hotspots (n = 7235 peaks; black line: mean; gray shading: SD). Predicted nucleosome positions are shown above. (B) Similar to A except for PRDM9Cst-dependent hotspots (n = 9306 peaks). (C) Fold enrichment of the PRDM9Dom2 motif over the PRDM9Cst motif at Prdm9Dom2-dependent hotspots in A. (D) Fold enrichment of the PRDM9Cst motif over the PRDM9Dom2 motif at PRDM9Cst-dependent hotspots in (B). (E) The PRDM9Dom2 motif identified in the NDRs of H3K4me3 peaks in A. (F) The PRDM9Cst motif.











