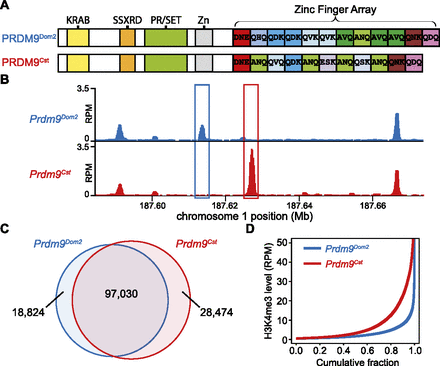
H3K4me3 ChIP-seq allows detection of PRDM9-dependent sites using co-isogenic strains of mice. (A) PRDM9Dom2 and PRDM9Cst differ both in the number of zinc finger domains and sequence-specific identity of individual domains. The known domain structure of PRDM9 includes an N-terminal KRAB domain (Krüppel-associated box), an SSXRD (SSX repression domain), a PR/SET domain (methyltransferase activity), an invariant zinc finger domain, and a C-terminal zinc finger array. The three most polymorphic positions in the individual zinc finger domains are indicated by their amino acid identity. (B) H3K4me3 coverage profile from a representative 100-kb window of chromosome 1. H3K4me3 peaks can be unique or shared between the two strains of mice (in all figures, the Prdm9Cst allele originates from the CAST/EiJ strain and is represented in red, while Prdm9Dom2, the endogenous allele for C57BL/6J, is represented in blue). (C) Venn diagram showing quantity and identity of H3K4me3 peaks. Prdm9Cst results in more unique H3K4me3-marked hotspots. (D) H3K4me3 signal for strain-specific peaks. Prdm9Cst hotspots have higher methylation levels than Prdm9Dom2 in the B6 background (red: Prdm9Cst; blue: Prdm9Dom2).











