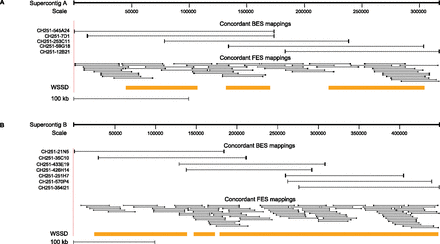
Figure 4.
Support for chimpanzee supercontig architecture from clone end mappings. Concordant BES and FES alignments confirm order and orientation of (A) distal and (B) proximal chimpanzee supercontig assemblies. One-hundred-twenty-five paired-end sequences that map with >99.8% sequence identity are depicted. Both analyses support high-quality assembly of these complex regions of the chimpanzee genome.











