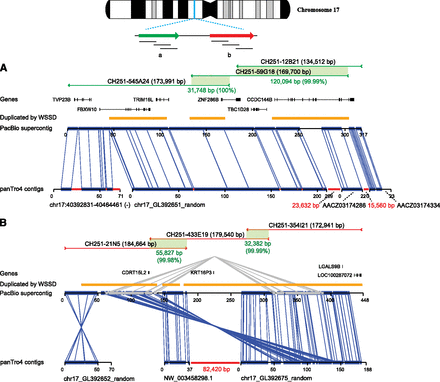
Upgrading a chimpanzee genomic region. Sequence and assembly of six large-insert clones (CH251) from two segmental duplication blocks (red and green) are aligned to their corresponding sequences from the 17p11.2 Smith-Magenis region of the chimpanzee reference assembly (panTro4). Clones were sequenced and assembled from the (A) distal and (B) proximal segmental duplication blocks. The PacBio assembly was compared with the corresponding working draft sequences from panTro4. The alignment identity of panTro4 contigs without gap sequence and the PacBio supercontigs is 94.69% over 525 kbp of aligned sequence. Thirty-one percent (241/766 kbp) of the chimpanzee sequence is missing within the working draft assembly. The average sequence identity for phred >30 bp from BAC end sequence (BES) mappings was 99.72% (16,174/16,220 high-quality bases) and 99.98% (156,955/156,991 high-quality bases) from fosmid end sequence (FES) mappings. Gaps in the panTro4 contigs are indicated in red. Gene annotations are shown based on a custom liftover from RefSeq annotations of GRCh37 in the corresponding regions of 17p11.2. The missing sequence corresponds to high-identity segmental duplications (orange bars represent segmental duplications predicted by whole-genome shotgun sequence detection or WSSD). The clone CH251-545A24 was previously sequenced with capillary sequencing (GenBank accession: AC183294).











