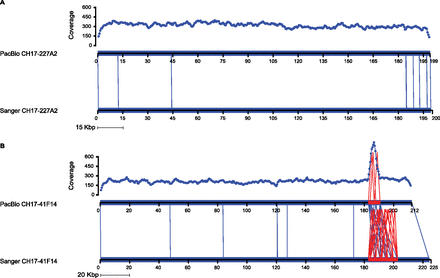
Concordant and discordant PacBio assemblies. (A) Alignment between PacBio (top) and Sanger (bottom) assemblies for CH17-227A2 using Miropeats (Parsons 1995) shows virtually no differences. Note the uniform sequence coverage between 200- and 300-fold. Mismatches/indels are indicated by vertical blue lines. (B) Alignment between PacBio and Sanger assemblies for clone CH17-41F14. A spike of increased sequence coverage across the internal repeat and the reduced complexity of the repeat compared with the Sanger assembly clearly define a collapse of a higher-order repeat from 20 to 12 kbp within the PacBio assembly. The uniformity of sequence coverage may be used as one indicator of potential misassembly.











