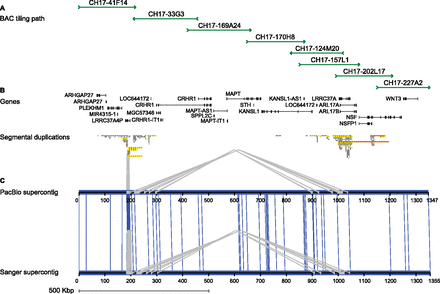
17q21.31 genomic target region. (A) Tiling path of eight large-insert BAC clones sequenced and assembled using both PacBio- and Sanger-based approaches. Clones were selected from a haploid complete hydatidiform mole source (CH17). (B) Gene annotation (RefSeq) and segmental duplication organization were obtained from GRCh37 using a custom liftover coordinate conversion tool that accounted for the difference in copy number between the mole haplotype and the reference. (C) Alignment of supercontigs built from the same eight clones using PacBio and Sanger assemblies. Sequence differences (vertical blue lines) and internal duplications (gray) are shown. The two supercontigs are 99.99% identical, excluding a collapsed higher-order repeat at the end of the PacBio assembly of CH17-41F14.











