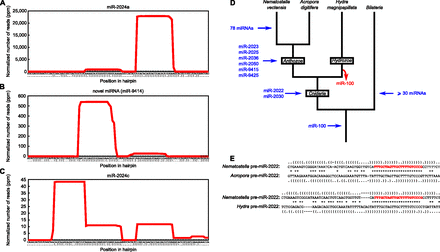
Discovery of novel Nematostella miRNAs. (A,B) Small RNA profiles along a pre-miRNA sequence (here exemplified by pre-miR-2024a and pre-nve-mir-9414). The x-axis indicates the nucleotide position along the hairpin sequence (predicted secondary structure is represented by dots and brackets, with dots indicating unpaired nucleotides and opening and closing brackets representing paired nucleotides). The y-axis indicates the number of reads covering each nucleotide in the pooled 18 deep-sequencing libraries. (C) A small RNA profile along endo-siRNA sequences (same conventions as in panels A and B). Several small RNAs (here exemplified by miR-2024c) were previously described as miRNAs, but the processing of their precursors into multiple small RNAs shows that these are actually siRNAs. (D) Proposed evolutionary scenario for the appearance of miRNA genes in the Nematostella vectensis lineage. The number of urbilaterian miRNAs (>30 in addition to miR-100) was estimated by comparing the known miRNA complement of a slowly evolving protostome (the annelid Capitella teleta) to the known miRNA complement of three deuterostomes (Strongylocentrotus purpuratus, Homo sapiens, and Tetraodon nigroviridis), requiring a perfectly conserved seed and an overall PHYLIP alignment score greater or equal than 0.3 (calculated by ClustalW). (E) Sequence and secondary structure alignment of Cnidarian pre-miR-2022. Mature miRNA sequences are shown in red. Conserved nucleotides are flagged with an asterisk. Secondary structures are represented by dots (for unpaired nucleotides) and brackets (for paired nucleotides). Dashes indicate alignment gaps.











