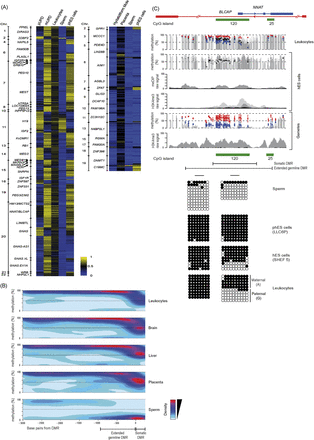
Methylation in gametes, hES cells, and somatic tissues. (A) Heat maps for Infinium probes mapping within all ubiquitous (left) and placental-specific (right) imprinted DMRs in sperm and phES cells reveal the germline acquisition of methylation. (B) Methylation contour plots from WGBS data sets for all maternally methylated DMRs reveal that the extent of the intermediately methylated regions associated with imprinted DMRs are extremely consistent between somatic tissues and significantly larger in sperm. (C) Methylation profiles at the NNAT DMR determined by WGBS, Infinium array, and meDIP-seq data sets in leukocytes, sperm, phES cells, and hES cells, along with the H3K4me3 ChIP-seq reads for hES cells and sperm. The gray and black dots in the second panel represent Infinium probe methylation in hES cell lines derived from six-cell blastomeres (Val10B) and blastocytes (SHEF5), respectively. The gametic WGBS methylation profile is derived from sperm, with Infinium probe methylation values for sperm and phES cells represented by blue and red dots. The graphic shows the extent of the differentially methylated regions in somatic tissues and between sperm and phES cells. The error bars associated with the Infinium array probes represent the standard deviation of the two sperm samples and four independent phES cell lines. The H3K4me3 ChIP-seq data is from sperm. The methylation profiles were confirmed using standard bisulfite PCR and sequencing.











