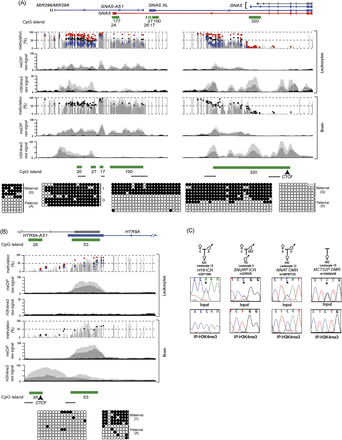
H3K4me3 chromatin profile and DNA methylation at imprinted loci. (A) Map of the human GNAS locus on chromosome 20 with the H3K4me3 and meDIP signatures in brain and leukocytes at the DMRs identified in the WGBS and Infinium array analysis. Infinium methylation values for normal leukocytes (black dots), with values for the genome-wide pUPD (blue) and mUPD (red) superimposed on the leukocyte WGBS track. Similarly, Infinium methylation values for two normal brain samples are shown as black and gray dots. The light and dark gray peaks in the meDIP and ChIP-seq panels represent two independent biological replicates compared to input (black peaks). The bars under the CpG islands, as identified in the UCSC Genome Browser, show the location of the bisulfite PCR amplicons. (B) The gradient DMR identified at the HTR5A promoter. The samples used for the WGBS, Infinium array, and ChIP are the same as in A. The independent methylation pattern on either side of the bidirectional promoter interval was confirmed using standard bisulfite PCR and sequencing. (C) Allelic ChIP for H3K4me3 reveals predominant enrichment of this histone modification on the unmethylated allele of the H19 ICR, SNURF ICR, NNAT, and MCTS2P DMRs. The asterisk (*) in the sequence traces shows the position of the polymorphic base.











