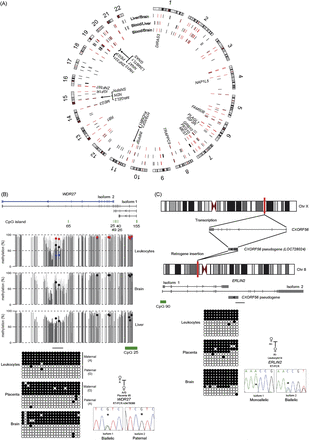
Identification and characterization of allelic methylation and expression of novel imprinted loci. Circular karyotype showing the position of common regions of intermediate methylation in the leukocyte, brain, and liver WGBS data sets, as identified using a 25 CpG sliding window approach (0.25 < mean ± 1.5 SD < 0.75). Red ticks represent sites of intermediate methylation common to all tissues, whereas black ticks identify those present in only one or two pairwise comparisons. The position of known imprinted DMRs are shown. (B) Identification of a novel maternally methylated DMR within the WDR27 locus by WGBS and Infinium array analysis. Vertical gray lines in the WGBS tracks represent the mean methylation value for individual CpG dinucleotides calculated from multiple data sets, with the light gray lines representing the mean + standard deviation. Infinium methylation values for normal tissues are represented by black dots, with values for the genome-wide UPDs (average pUPD in blue and mUPD in red) superimposed on the leukocyte methylation track. The DMR was confirmed using standard bisulfite PCR on heterozygous DNA samples and orchestrates paternal expression of WDR27 isoform 2. The asterisk (*) in the sequence traces shows the position of the polymorphic base. (C) Imprinting of ERLIN2 isoform 1 in leukocytes as a consequence of the retrotransposition of the X chromosome-derived CXorf56 pseudogene into the locus.











