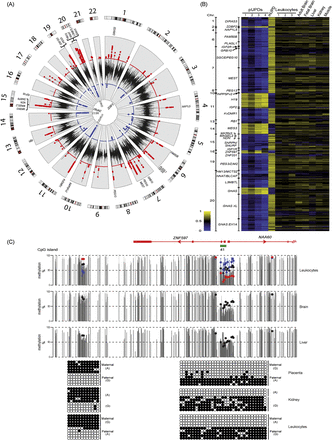
Identification of known imprinted DMRs on the Infinium array platform. (A) Circular karyotype showing the difference of methylation for three consecutive probes for reciprocal UPD leukocyte samples. Red dots indicate a minimal difference of 0.3 in Infinium probe β-values (>30% absolute methylation value) for regions with maternal methylation, and blue dots indicate the same for paternal methylation. Known DMRs are indicated. (B) Heat map of the Infinium probes located within known imprinted DMRs in reciprocal genome-wide UPD samples and various somatic tissues. (C) WGBS and Infinium array methylation profiles of the ZNF597 locus with bisulfite PCR confirmation of the novel maternally methylated DMR and its position in relation to the somatic paternally methylated promoter region. Vertical gray lines in the WGBS tracks represent the mean methylation value for individual CpG dinucleotides calculated from multiple data sets, with the light gray lines representing the mean + standard deviation. Infinium methylation values for normal tissues are represented by black dots, with values for the genome-wide UPDs (average pUPD in blue and mUPD in red) superimposed on the leukocyte methylation track. The error bars associated with the Infinium array probes represent the standard deviation of multiple biological samples. The PCR confirmation in placenta, kidney, and leukocyte-derived DNA was performed on heterozygous samples. Each circle represents a single CpG dinucleotide on a DNA strand. (●) Methylated cytosine, (○) unmethylated cytosine. Each row corresponds to an individual cloned sequence.











